The structure of DNA (e.g., gene insert, a recombinant plasmid or entire genome) can be analysed by determining the nucleotide sequences. In molecular cloning, the information of nucleotide sequences are essential. In 1965, Robert Holley and his research group at Cornell University completely sequenced nucleotides of tRNAala (tRNA for yeast alanine). In 1977, the following two methods were developed.
Allan Maxam and Walter Gilbert developed a chemical method of DNA sequencing. In this method, end-labelled DNA is subjected to base specific cleavage reaction before gel separation. In routine sequencing of DNA this method is not commonly followed. In the same year (1977) Frederick Sanger and co-workers developed an enzymatic method of DNA sequencing. It is also called dideoxynucleotide chain termination method because dideoxynucleotides are used as chain terminator to produce a ladder of molecules.
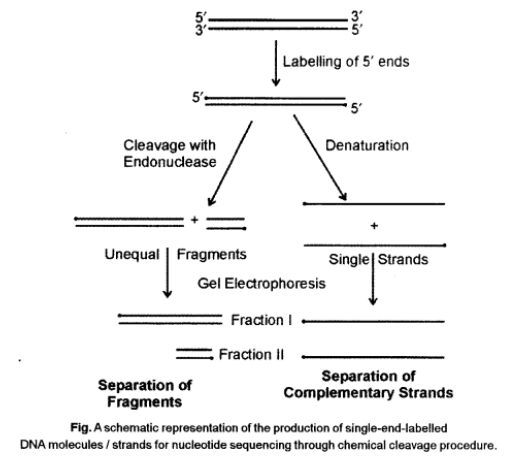
Maxam and Gilbert’s Chemical Degradation Method
In this method, DNA sequencing involves the following steps :
• Labelling of 3 ’ends of DNA with isotopic phosphorus (32P).
• Separation of two strands labelled at 3 ’ ends.
• Separation of mixture in four sets, each treated with a different reagent which can degrade only G or C, or A and G or T and C.
• Electrophoretic separation of each sample in four different gel.
• Autoradiography of gels and determination of the sequence from position of bands in four lanes of gel.