Restriction enzymes are bacterial proteins that have the ability, to cut both the strands of the DNA molecule at a specific nucleotide sequence. There are hundreds of these restriction enzymes and each can cut the DNA at a specific point and the resulting DNA fragments are all of different lengths. Restriction enzymes can either produce sticky ends or blunt ends.
EcoRI enzyme binds to a region of having specific palindromic’ sequence (where two strands are identical when both are read in the same polarity i.e., in 5’→ 3’ direction). The length of this region is 6 base pairs i.e., hexanucleotide palindrome. It cuts between G and A residues of each strand and produces two single stranded complementary cut ends which are asymmetrical having 5’ overhangs of 4 nucleotides. These ends are called sticky ends or cohesive ends. Because nucleotide bases of this region can pair and stick the DNA fragments again as given below :}

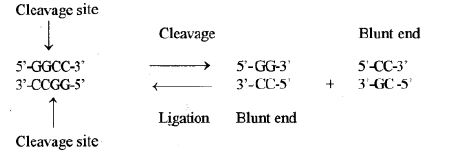
On the other hand, there are some other Type II restriction enzymes which cleave both strands of DNA at the same base pairs but in the centre of recognition sequence, and results in DNA fragments with blunt ends or flush ends.
For example
Hae111 (isolated from Haemophilus aegypticus, the order of enzyme III), four nucleotide long palindromic sequence and cuts symmetrically the both DNA strands and forms blunt ends as below: